Frequently Asked Questions (FAQ)
Access and License
Are the dataset submissions open to all researchers?
- Answer: The repository in Open Access to all researchers. Viewing the data is possible without limitation, the download of datasets is only possible if the users are registered (to avoid misuse) but all interested people can register. The submission of datasets requires registration (for further correspondence and curation requests if applicable) but is open to all scientists.
Under which kind of license are the data of the Chemotion-Repository available?
- Answer: Data are available under CC by SA 4.0, CC by or CC 0 license. Authors can also publish contents under public domain mark.
Does the Chemotion-Repository charge access fees or subscription fees?
- Answer: No. Viewing the data is possible without limitation (Open Access), only the download of datasets is limited to registered users (to avoid misuse). All interested people can register easily and without any costs or disadvantages.
How is data accessible if one set is an embargo?
- Answer: The embargo can only be released by the submitter of the datasets. Only if all data that belong to the embargo bundle are complete and reviewed, the release is possible. Please see also the Usage Pages for further information.
Data Persistence
Does the repository assign a stable persistent identifier (PID) for each dataset at publication, such as a digital object identifier (DOI)?
- Answer: The Chemotion-Repository assigns a DOI for each dataset (and molecules, reactions) via DataCite. The name used in the Publisher namespace is “Chemotion.net”. The Chemotion-Repository is assigned to the DOI string 10.14272.
Is there a long-term data management plan (including funding) to ensure that datasets are maintained for the foreseeable future.
- Answer: The repo will be part of the initiative Science Data Center of the federal state of Baden-Württemberg (https://mwk.baden-wuerttemberg.de/de/service/presse/pressemitteilung/pid/vier-science-data-centers-in-baden-wuerttemberg/) and is planned for the national research data infrastructure in Germany. A long term storage and archival plan is developed with the Scientific Center for Computing under these umbrellas.
Are researchers able to modify or remove datasets after publication?
- Answer: Only the submission of a new version is possible. A new version can be sent but the old one will be still visible/accessible.
Do you guarantee persistent access to datasets, and for how long?
- Answer: For at least 10 years.
How long has your resource been available to the community?
- Answer: Since 2014 (with interruption due to major updates/rework in 2016-2018).
Data Processing
Is there is any curation support for researchers uploading their datasets? If so, please describe this briefly.
- Answer: The data is curated manually by peer reviewing supported by an automated analysis of NMR data. The analysis of NMR data is compared automatically with the expected signals (according to the formula of the compounds provided).
Do you capture any metadata about hosted datasets in a standardized way? If so, please state which metadata formats are used.
- Answer: Metadata support is provided for information like analysis types and reaction names. The metadata can be selected via a dropdown menus (supports search) that provides the embedded ontobee ontologies CHEMINF and CHMO http://www.ontobee.org/. We use the metadata format of DataCite for DOI submissions.
Community
How large is your current user base?
- Answer: Users with registration: 650. To access the data, no registration is necessary. The access to data is not tracked.
How many datasets are currently hosted by the repository?
- Answer: Information on the data availability is given at the landing page of the repository, e.g. 1546 published reactions (+ 1804 reactions under embargo or in reviewing status [date: 12.03.2023])
Are there examples that demonstrate the acceptance within the relevant research community?
- Answer: Examples showing acceptance by reviewers and strong integration of the repository data to a publisher’s website are given below. Other examples can be found in the repository as well as from links to the datasets from relevant publications.
- The first published article with full data deposition in Chemotion-Repository with referencing of all datasets in the SI. All data was published in the repository before the final acceptance of the publication: N. Jung, S. Grässle, D. S. Lütjohann, S. Bräse, Org. Lett. 2014, 16, 1036.
- Another example, where the publisher developed novel options for the listing of research data based on the functions of the repository (indexing per RInCHI and link to Chemotion-Repository): Y.-C. Huang, A. Nguyen, S. Vanderheiden, S. Gräßle, N. Jung, S. Bräse, Beilstein J. Org. Chem. 2018, 14, 515.
- The relevance in materials sciences could be demonstrated by a recently accepted manuscript: Synthesis of Functionalized Azobiphenyl- and Azoterphenyl- Ditopic Linkers: Modular Building Blocks for Photoresponsive Smart Materials, S. Grosjean, P. Hodapp, Z. Hassan, C. Wöll, M. Nieger, S. Bräse, ChemistryOpen 2019, 8, 743.
Are there any entries in other databases for the Chemotion-Repository?
- Answer: Chemotion is listed in
- re3data: https://www.re3data.org/repository/r3d100010748
- FAIRsharing.org: https://fairsharing.org/biodbcore-001268
- RiSources: https://risources.dfg.de/detail/RI_00351_en.html
Is there a repository Twitter handle or similar activities?
- Answer: Hints and demos for the repository and the ELN can be found on this YouTube video.
Has the repository undergone WDS (World Data Systems), DSA (Data Seal of Approval) or CTS (Core Trust Seal) accreditation?
- Answer: No, but we will apply for a Core Trust Seal soon.
Capabilities
What type of experimental data can be hosted by the repository? (If the repository only accepts specific file formats please state what these are.)
- Answer: There is no limitation with respect to file formats. Open File formats are preferred and special visualization tools for e.g. JCAMP spectra exist.
What is the maximum file size that can be handled by the repository?
- Answer: No limitation so far but we might limit it to 50 MB in future.
Are there any limitations to the amount of data that an individual is able to upload?
- Answer: So far not. We will limit the size if misuse is detected. The data to be disclosed are reviewed and misuse will be detected.
Does the repository have the facility to provide controlled access to sensitive data?
- Answer: Users have the option to collect data on their private account (1) without disclosure or (2) to disclose the data with an embargo. If the embargo option is selected, the data will, after reviewing, be available only for the users and external reviewers (by mail notification if desired by the user) and release will take place after additional confirmation by the user.
Is the repository able to facilitate confidential peer review of hosted datasets? If yes, please briefly describe the workflow for reviewing hosted datasets, including how reviewers may access data which are not public at the time of review.
- Answer: Reviewing first takes place by an automated quick check of most common data like NMR data (counting of signals necessary and analyzed) and in addition by a peer reviewing by the repository owners. Reviewing workflow includes comment functions for the datasets that allows to reject data and to ask for revision (mediated via the repository UI and per email for notifications). If the data is submitted to the repo with embargo, the user has the option to provide access to the data to single external reviewers (access given by limited accounts, provided by mail).
How does the infrastructure of Chemotion repository look like?
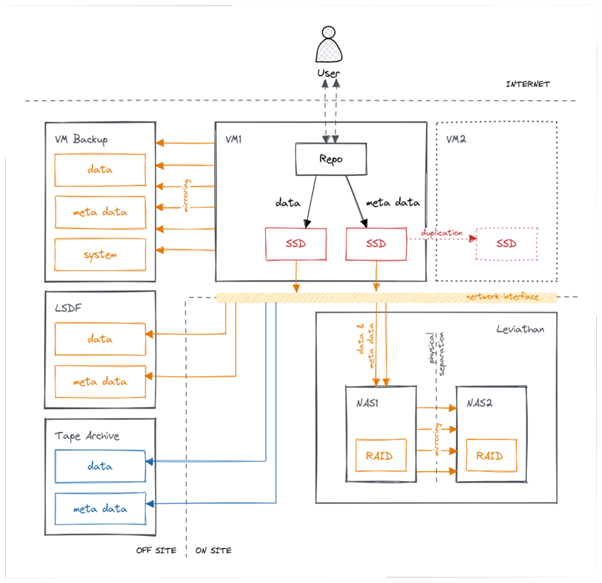
Linking
How can I link hosted datasets to relevant articles after publication?
-
Answer: The repository submissions can be assigned to a doi or a reference added to single data submissions or a collection of data submissions.
In addition, the repository provides a function to retrieve a virtual DOI for the dataset even if the data is not disclosed yet. This allows to give the correct doi already in the supporting information of a publication even if the dataset is not available to the public yet. This allows a direct link of publication and dataset in the SI (for example). Please see or „How To topics“.
Are there some examples of how to present the doi of datasets in articles or the SI?
-
Answer: For the link of data to the repository, please see the supporting information of the selected examples:
Example 1: link to publication:
- https://www.beilstein-journals.org/bjoc/articles/15/114/downloads Link to Datasets (examples):
- https://www.chemotion-repository.net/home/publications/molecules/1408
- https://www.chemotion-repository.net/home/publications/molecules/1345
- https://www.chemotion-repository.net/home/publications/molecules/1339
Example 2: link to publication:
Link to Datasets (examples):